Starting a Research Group
Tim Witney
It’s been a while since I last posted on here; over 3 years in fact. The original aim of this blog was to try and summarise some of our recently-published work without the scientific jargon that not only makes these papers incomprehensible to most people, but has the added disadvantage of putting them off what might (or might not!) be an important topic. This was exemplified recently when one of the papers from our lab was published in the scientific Journal Cancer Research. I asked my family how many words into the abstract they could get before getting lost. I lost them all on the third word - ‘endogenous’. This problem was magnified by the fact that the title made no sense to them either. So far, not so good. So I’m going to try a little harder to find the time to post here again in order to explain the science that we do and why I think it’s important.
Why haven’t you posted anything for ages I hear no one ask? Well, for the last ~5 years I have been been busy starting my own research Group. I say my own, but it’s not my own as it’s jointly owned by the very talented scientists that I get to work with every day (see pic). Together, we are part of a far larger Department at King’s where collaboration is deeply ingrained and the boundaries of each Group are blurred. But it’s taken me nearly 5 years to get to a point where I can reflect on this sometimes difficult process, which has seen me move Universities and start all over again. I thought I’d write about what it’s like to start a research Group, the errors that I have made, and the joys of being able to perform the science that really interests me.
The five stages of grief
Don’t worry, this story has a happy ending! But starting a Group is not easy. Up until this point we have been trained to be very good scientists but no thought has been put into what it takes to be a good manager, how to handle finances or handle high-level project management. So doing these things for the first time can often feel like a big shock to the system and has probably been the hardest thing I’ve done in my career to-date. I forgot who told me this, but they compared starting your own lab to the five stages of grief, which on refection is a great analogy. It’s reproduced below. I should stress that this is in no-way the process that everyone goes through, but it resonated with me. Of course it goes without saying that this is just a way of illustrating the different emotions I went though over the past five years and in no way can be compared to the tragedy of losing someone important to you.
Denial
It all starts with money. Actually, that’s not strictly true - it starts with an idea, but that idea will die unless you can get a research grant to fund people and materials to enact that idea. I was fortunate enough to be awarded a Wellcome Trust & Royal Society Sir Henry Dale Fellowship that enabled me to do just that. It’s hard to describe the joy of receiving the news that my Fellowship had been funded. But after that initial euphoria dies down you’re left with the task of having to hire for the very first time, find space to perform experiments, get quotes for everything from the most important piece of equipment to a basic fridge. The list is long; sometimes exhaustingly so, which can lead to avoidance of the issues, procrastination (see twitter and this website) and an overwhelming sense of the magnitude of the task. Luckily I had thought a lot about what I would like my Group to look like, but there’s no substitution for actually going through this experience yourself. In many way it’s like becoming a parent for the first time - no matter how many people you talk to on the subject, nothing prepares you for the real thing.
2. Anger
Anger is probably a little extreme, but once you’ve got everything together and you’re ready to go you’d think that this would be it and everything would fall into place. Unfortunately, everything takes longer than you first thought. From hiring the right person (sometimes this means having to readvertise), finding lab space that was initially promised to you which has now mysteriously disappeared, to the dawning realisation that politics plays a prominent role in science and you’re the bottom of the rung. It’s not anger, but more of a frustrating process of having to get all your ducks in a row. One of the most annoying things that’s immediately obvious is the fact that suddenly you have to prove yourself all over again. As if the past maybe 8 years or so was a combination of luck and your previous PI’s genius. Which I can’t rule out in my case!
3. Bargaining
You might have some equipment and if you’re lucky, a postdoc or grad student to micromanage (sorry, I mean mentor…), but quickly I realised that a great idea, willing hands and some pipettes is not enough and you really need some outside help to sustain you and your research in the early days. There are many colleagues over the last few years that have helped me with no expectation of the help being repaid, be it reading drafts of grants, acting as a sounding board, or having the patience to teach your student a new technique. However, further afield, forget it if you think people will magically know either who you are or will want to collaborate straight away. You have no track record as a PI, so it’s often seen as a risk. People are not knocking down your door to work with you. So how to get around this problem in the early days? Often it’s a case of bargaining - finding areas of mutual interest, supporting research that’s not directly related to your own programme and frequently exchanging important data and analysis for a name on the paper that is yet to be written.
4. Depression
Which is where the shock hits. Having pretty consistently published at least one first author paper a year during my postdocs, the reality is that even with many more people working on the same idea or project, it takes over double the amount of time to get your first paper published. In fact, it took us more than three years before we started sending our paper out for rejection. I mean, publication. This first paper is often your baby, so you know exactly what form it will take. But there’s a lot of training to do. And now you’re needed to teach, be on interview panels, speak at conferences, complete all the admin the U.K.’s University system seems to thrive on. Things take their time in the lab. You might not have the same resources you were used to either. This is all against the background of some sort of ticking clock when you know you have to either deliver or be out of a job, ~10-15 years after starting and with no hope of surviving in the ‘real world’. I was really feeling the pressure three years in and I think not only I, but my Team suffered because of the worry that surrounded the fixed-term contract that I was on. This is maybe a topic for another day, but Universities really need to look hard at how they support their Research Fellows. Put your money where your mouth is and stop taking advantage of highly-skilled Young Investigators. Sorry, I’ve gone back to step 2. Back on topic, once the first paper from your Group is ready, there is so much excitement surrounding the paper and the endless possibilities. Why wouldn’t Nature take it?! I mean, we’ve all seen worse published there. You may be lucky, but we aimed high and were persistent. So were the desk rejections. The few months before our first work was published in Cancer Research was a tough time.
5. Acceptance
But then things start clicking into place. You find your rhythm managing people, some grants are funded, and a lot aren’t. But you start getting used to rejection as part of the everyday way of things. It’s a necessary, but unfortunate side product of running your own Group. From one idea new data spawns many more; made all the more joyous when these come from other members of the Team. Publishing certainly helps the confidence and the realisation that running a research lab is a good fit for you. Acceptance that you are learning and won’t get everything right all the time also comes. I used to spend a lot of time worrying when I got things wrong, but now I try and spend the time planning for that difficult conversation or learning from past mistakes. Acceptance is also about admitting when things aren’t quite right. This realisation was the catalyst for me moving from UCL down the road to King’s.
There are some beautiful moments that come from running a lab. Some of my highlights include seeing my former PhD supervisor frantically scribbling down notes during one of my talks, the excitement of my PhD student hearing her project described for the first time, and being invited by your long-time friends to see their labs and speak at their seminars. It’s not for everyone, but it’s certainly for me.


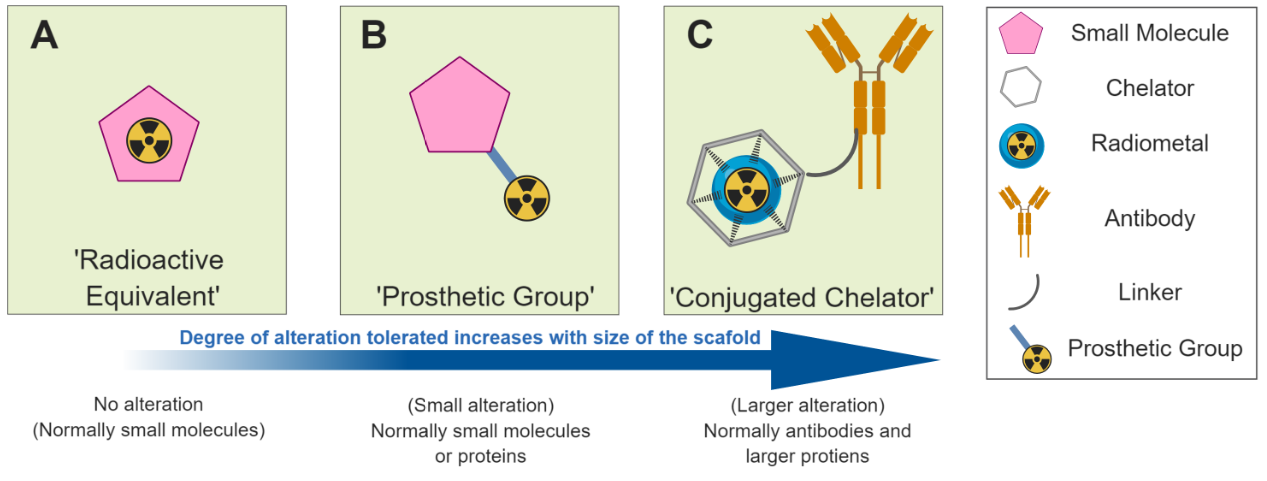
![Figure 2. Different radiolabelled scaffolds and their applications. A) [18F]FDG is transported in to cells where it accumulates due to metabolic trapping subsequent to phosphorylation by hexokinase. B) A radiolabelled drug molecule crosses the cellu…](https://images.squarespace-cdn.com/content/v1/53db178ce4b07b7551af4b99/1565002879783-O87DX5OS8O4C0M31UQZ3/Figure+2.PNG)